
Text to SBGN
For an example we chose the paper by Song and co-authors “Parkin promotes proteasomal degradation of p62: implication of selective vulnerability of neuronal cells in the pathogenesis of Parkinson’s disease” published in Protein Cell in 2016 (PMID: 26746706).
The complete abstract text from PubMed for this paper is here:
“Mutations or inactivation of parkin, an E3 ubiquitin ligase, are associated with familial form or sporadic Parkinson’s disease (PD), respectively, which manifested with the selective vulnerability of neuronal cells in substantia nigra (SN) and striatum (STR) regions. However, the underlying molecular mechanism linking parkin with the etiology of PD remains elusive. Here we report that p62, a critical regulator for protein quality control, inclusion body formation, selective autophagy and diverse signaling pathways, is a new substrate of parkin. P62 levels were increased in the SN and STR regions, but not in other brain regions in parkin knockout mice. Parkin directly interacts with and ubiquitinates p62 at the K13 to promote proteasomal degradation of p62 even in the absence of ATG5. Pathogenic mutations, knockdown of parkin or mutation of p62 at K13 prevented the degradation of p62. We further showed that parkin deficiency mice have pronounced loss of tyrosine hydroxylase positive neurons and have worse performance in motor test when treated with 6-hydroxydopamine hydrochloride in aged mice. These results suggest that, in addition to their critical role in regulating autophagy, p62 are subjected to parkin mediated proteasomal degradation and implicate that the dysregulation of parkin/p62 axis may involve in the selective vulnerability of neuronal cells during the onset of PD pathogenesis.”
After omitting some words and keeping it close to the original text, the key points for our diagram are these:
- Mutations of parkin are associated with Parkinson’s disease (PD) and selective vulnerability of neuronal cells in substantia nigra (SN) and striatum (STR).
- Here we report that p62 is a new substrate of parkin. p62 levels were increased in the SN and STR regions in parkin knockout mice.
- Parkin directly interacts with and ubiquitinates p62 at the K13 to promote proteasomal degradation of p62.
- We further showed that parkin deficiency mice have pronounced loss of tyrosine hydroxylase positive neurons.
- These results suggest that p62 are subjected to parkin mediated proteasomal degradation and implicate that the dysregulation of the parkin/p62 axis may be involved in the selective vulnerability of neuronal cells during the onset of PD pathogenesis.
Point #3 gives enough information for drawing a detailed Process Description diagram in CellDesigner (Figure 1). Other points provide information about the cell type (neuron) and the context (Parkinson’s disease). Point #4 can be potentially explored to link these mechanisms to tyrosine hydroxylase (TH, UniProt:P07101, HGNC:11782) in the context of Parkinson’s disease but for that more information is required.
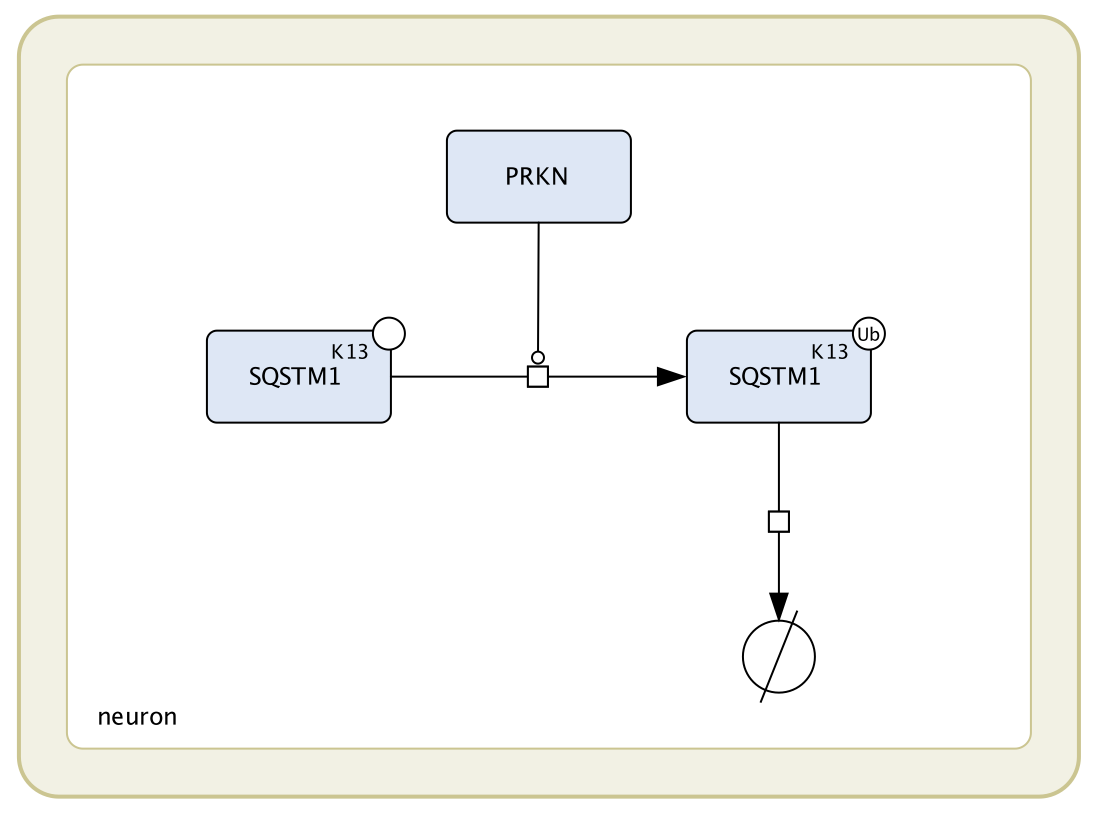
Figure 1. The process of SQSTM1 ubiquitination at lysine 13 activated by PRKN, followed by the process of degradation of the ubiquitinated SQSTM1.
Please note that for the diagram we use the official HUGO Gene Nomenclature Committee (HGNC) names for proteins. For that “parkin” is replaced with the official “PRKN” (UniProt:O60260, HGNC:8607, previously PARK2) and “p62” is replaced with the official “SQSTM1” (UniProt:Q13501, HGNC:11280).